|
|
||||
|
|
||||
|
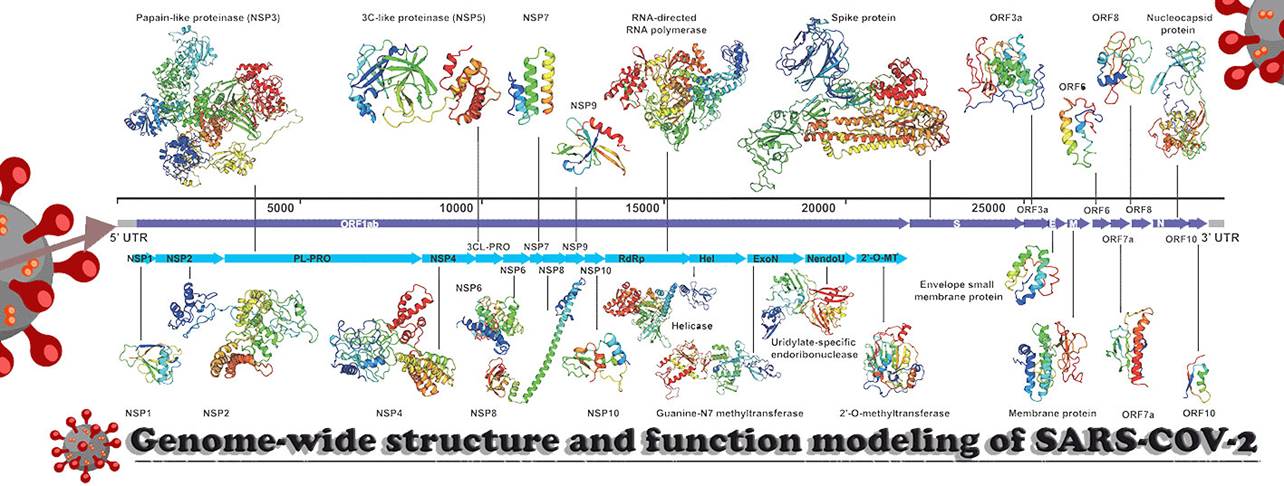
SARS-CoV-2 protein components. Summary and comments of PDB
entries |
||||
|
Protein component |
Function |
PDB ID and link |
Resolution method |
Ligand |
|
Nsp1 |
It suppresses host gene expression by
ribosome association. It blocks the innate immune responses that would
otherwise facilitate clearance of the infection |
Electron Microscopy |
human 40S ribosomal subunit |
|
|
Electron Microscopy |
human
CCDC124-80S-EBP1 ribosome complex
|
|||
|
Electron Microscopy |
human
CCDC124-80S-eERF1 ribosome complex
|
|||
|
Electron Microscopy |
human LYAR-80S ribosome complex |
|||
|
Electron Microscopy |
pre-40S-like ribosome complex |
|||
|
Electron Microscopy |
pre-40S-like ribosome complex - state 2 |
|||
|
Electron Microscopy |
human 43S preinitiation ribosome complex - state 2 |
|||
|
Nsp2 |
Nsp2, along with nsp1, are involved in
suppressing gene expression |
- |
|
|
|
Nsp3 |
Involved in viral replication. The Nsp3-4-6
complex regulates the replication site by recruiting the replicase proteins to
the host membrane. Nsp3, Nsp4 and Nsp6 assemble at the endoplasmic reticulum,
driving its modification and formation of the DMV (double membrane vesicles)
Papain-like protease domain |
X-Ray Diffraction |
AMP (ADENOSINE MONOPHOSPHATE) |
|
|
X-Ray Diffraction |
peptide
inhibitor VIR250
|
|||
|
X-Ray Diffraction |
|
|||
|
X-Ray Diffraction |
|
|||
|
X-Ray Diffraction |
ADP ribose |
|||
|
X-Ray Diffraction |
MES, 2-(N-MORPHOLINO)-ETHANESULFONIC ACID |
|||
|
X-Ray Diffraction |
|
|||
|
X-Ray Diffraction |
|
|||
|
X-Ray Diffraction |
|
|||
|
X-Ray Diffraction |
peptide inhibitor VIR251 |
|||
|
X-Ray Diffraction |
P3221 space group |
|||
|
X-Ray Diffraction |
ubiquitin
propargylamide
|
|||
|
X-Ray Diffraction |
SG15 C-terminal domain propargylamide |
|||
|
X-Ray Diffraction |
|
|||
|
X-Ray Diffraction |
HEPES
|
|||
|
X-Ray Diffraction |
ADP-ribose
|
|||
|
X-Ray Diffraction |
MES,
2-(N-MORPHOLINO)-ETHANESULFONIC ACID
|
|||
|
Nsp4 |
Nsp3-4-6 complex for endoplasmic reticulum
modification and formation of DMVs |
- |
|
|
|
Nsp5 |
Main protease |
X-Ray Diffraction |
inhibitor N3 |
|
|
X-Ray Diffraction |
inhibitor 11a |
|||
|
X-Ray Diffraction |
|
|||
|
X-Ray Diffraction |
inhibitor 11b |
|||
|
X-Ray Diffraction |
novel inhibitor |
|||
|
|
|
|||
|
X-Ray Diffraction |
non-covalent inhibitor X77 |
|||
|
X-Ray Diffraction |
inhibitor GC-376 |
|||
|
X-Ray Diffraction |
Leupeptin |
|||
|
X-Ray Diffraction |
|
|||
|
X-Ray Diffraction |
Oxidized C145 (sulfinic acid cysteine) |
|||
|
X-Ray Diffraction |
α-ketoamide inhibitor. |
|||
|
X-Ray Diffraction |
|
|||
|
X-Ray Diffraction |
α-ketoamide inhibitor. |
|||
|
X-Ray Diffraction |
|
|||
|
X-Ray Diffraction |
|
|||
|
X-Ray Diffraction |
2-Methyl-1-tetralone |
|||
|
X-Ray Diffraction |
mISG15 |
|||
|
X-Ray Diffraction |
AZD6482 |
|||
|
X-Ray Diffraction |
biotin-PEG(4)-Abu-Tle-Leu-Gln-vinylsulfone |
|||
|
X-Ray Diffraction |
Boceprevir |
|||
|
X-Ray Diffraction |
Feline coronavirus drug |
|||
|
X-Ray Diffraction |
Feline coronavirus drug |
|||
|
X-Ray Diffraction |
Feline coronavirus drug |
|||
|
X-Ray Diffraction |
UAW241 |
|||
|
X-Ray Diffraction |
|
|||
|
X-Ray Diffraction |
|
|||
|
X-Ray Diffraction |
|
|||
|
X-Ray Diffraction |
UAW246 |
|||
|
X-Ray Diffraction |
UAW247 |
|||
|
X-Ray Diffraction |
UAW248 |
|||
|
X-Ray Diffraction |
Ketone-Based Covalent Inhibitors |
|||
|
6XHU |
X-Ray Diffraction |
|
||
|
6XFN |
X-Ray Diffraction |
UAW243 |
||
|
X-Ray Diffraction |
oxidized Cys145 |
|||
|
X-Ray Diffraction |
inhibitor 7j |
|||
|
X-Ray Diffraction |
Leupeptin |
|||
|
7B83 |
X-Ray Diffraction |
pyrithione zinc |
||
|
X-Ray Diffraction |
Boceprevir |
|||
|
X-Ray Diffraction |
Inhibitor N3 |
|||
|
X-Ray Diffraction |
carmofur |
|||
|
X-Ray Diffraction |
TG-0203770 |
|||
|
X-Ray Diffraction |
TG-0205221 |
|||
|
X-Ray Diffraction |
GC376 |
|||
|
X-Ray Diffraction |
GC376 |
|||
|
Nsp6 |
Nsp3-4-6 complex for endoplasmic reticulum
modification and formation of DMVs |
- |
|
|
|
Nsp7-Nsp8-Nsp12 Complex (RNA-dependent RNA polymerase) (RdRp) |
Part of the replication-transcription complex, involved in
virus replication |
Electron Microscopy |
nsp7-nsp8c complex |
|
|
Electron Microscopy |
|
|||
|
Electron Microscopy |
nsp7-nsp8c complex |
|||
|
Electron Microscopy |
nsp7-nsp8c complex |
|||
|
Electron Microscopy |
nsp7-nsp8c complex |
|||
|
Electron Microscopy |
nsp7, nsp8, nsp12 bound to nsp13 |
|||
|
Electron Microscopy |
nsp7-nsp8c complex |
|||
|
Electron Microscopy |
nsp7-nsp8c complex |
|||
|
Electron Microscopy |
nsp7-nsp8c
complex
|
|||
|
Electron Microscopy |
|
|||
|
7AAP |
Electron Microscopy |
primer dsRNA and favipiravir-RTP |
||
|
Electron Microscopy |
|
|||
|
Electron Microscopy |
|
|||
|
Electron Microscopy |
template-primer RNA and triphosphate form of Remdesivir(RTP) |
|||
|
Electron Microscopy |
|
|||
|
Electron Microscopy |
nsp7-nsp8c complex |
|||
|
Electron Microscopy |
nsp7-nsp8c complex |
|||
|
Nsp9 |
RNA replicase |
X-Ray Diffraction |
|
|
|
X-Ray Diffraction |
|
|||
|
X-Ray Diffraction |
|
|||
|
Nsp10-Nsp16 |
Nsp10 plays multiple roles in RNA capping |
X-Ray Diffraction |
nsp16-nsp10 with SAM (S-ADENOSYLMETHIONINE) |
|
|
X-Ray Diffraction |
nsp16-nsp10 complex |
|||
|
X-Ray Diffraction |
nsp16-nsp10 complex |
|||
|
X-Ray Diffraction |
nsp16-nsp10 complex |
|||
|
X-Ray Diffraction |
nsp16-nsp10 with
S-Adenosyl-L-Homocysteine |
|||
|
X-Ray Diffraction |
nsp16-nsp10 7-methyl-GpppA and S-adenosyl-L-homocysteine |
|||
|
X-Ray Diffraction |
nsp16-nsp10 with RNA cap analogue (m7GpppA) and
S-adenosylmethionine |
|||
|
X-Ray Diffraction |
nsp10-nsp16 with Sinefungin |
|||
|
X-Ray Diffraction |
nsp16-nsp10 with 7-methyl-GpppA,
S-adenosyl-L-homocysteine and Sulfates |
|||
|
X-Ray Diffraction |
nsp16-nsp10 with 7-methyl-GpppA and S-Adenosylmethionine |
|||
|
X-Ray Diffraction |
nsp10-nsp16 with Sinefungin |
|||
|
X-Ray Diffraction |
nsp10 |
|||
|
X-Ray Diffraction |
nsp16-nsp10 complex |
|||
|
X-Ray Diffraction |
nsp16-nsp10 with SAM |
|||
|
X-Ray Diffraction |
nsp16-nsp10 with SAM |
|||
|
Nsp11 |
|
- |
|
|
|
Nsp13 |
|
Electron Microscopy |
nsp7, nsp8, nsp12 bound to nsp13 |
|
|
Nsp14 |
|
- |
|
|
|
Nsp15 |
RNA endonuclease |
X-Ray Diffraction |
|
|
|
X-Ray Diffraction |
Citrate |
|||
|
X-Ray Diffraction |
Uridine-5'-Monophosphate |
|||
|
X-Ray Diffraction |
Tipiracil |
|||
|
X-Ray Diffraction |
Nucleotide GpU |
|||
|
X-Ray Diffraction |
|
|||
|
S |
Spike (S) glycoprotein |
X-Ray Diffraction |
S2 subunit |
|
|
X-Ray Diffraction |
Receptor Binding Domain (RBD) of S with ACE2 |
|||
|
X-Ray Diffraction |
S2 subunit |
|||
|
X-Ray Diffraction |
RBD with ACE2 |
|||
|
Electron Microscopy |
RBD with ACE2-B0AT1 complex |
|||
|
X-Ray Diffraction |
S2 subunit |
|||
|
X-Ray Diffraction |
RBD with EY6A Fab |
|||
|
Electron Microscopy |
|
|||
|
Electron Microscopy |
|
|||
|
Electron Microscopy |
|
|||
|
X-Ray Diffraction |
RBD with ACE2 |
|||
|
Electron Microscopy |
|
|||
|
Electron Microscopy |
|
|||
|
X-Ray Diffraction |
RBD with CR3022 |
|||
|
Electron Microscopy |
S309 neutralizing antibody Fab fragment |
|||
|
Electron Microscopy |
S309 neutralizing antibody Fab fragment |
|||
|
Electron Microscopy |
|
|||
|
Electron Microscopy |
|
|||
|
Electron Microscopy |
|
|||
|
Electron Microscopy |
|
|||
|
Electron Microscopy |
|
|||
|
X-Ray Diffraction |
RBD with antibody CC12.1 |
|||
|
X-Ray Diffraction |
RBD with antibodies CC12.1 and CR3022 |
|||
|
X-Ray Diffraction |
RBD with antibody CC12.3 |
|||
|
Electron Microscopy |
C105 neutralizing antibody Fab fragment (state 1) |
|||
|
Electron Microscopy |
C105 neutralizing antibody Fab fragment (state 2) |
|||
|
Electron Microscopy |
RBD with the Fab fragments of two neutralizing antibodies |
|||
|
X-Ray Diffraction |
RBD with CV30 Fab |
|||
|
Electron Microscopy |
|
|||
|
Electron Microscopy |
|
|||
|
Electron Microscopy |
|
|||
|
X-Ray Diffraction |
RBD with CR3022 Fab |
|||
|
X-Ray Diffraction |
RBD with CR3022 Fab |
|||
|
Electron Microscopy |
S1 unit with CR3022 Fab |
|||
|
X-Ray Diffraction |
RBD with nanobody H11-D4 |
|||
|
Electron Microscopy |
|
|||
|
Electron Microscopy |
|
|||
|
X-Ray Diffraction |
RBD with an antibody |
|||
|
Electron Microscopy |
BD23-Fab |
|||
|
X-Ray Diffraction |
RBD with CB6 antibody |
|||
|
Electron Microscopy |
4A8 |
|||
|
Electron Microscopy |
|
|||
|
Electron Microscopy |
|
|||
|
Electron Microscopy |
Trimerized VH binder |
|||
|
Electron Microscopy |
SARS-CoV-2
Omicron spike protein
|
|||
|
Electron Microscopy |
SARS-CoV-2
Omicron spike protein in complex with human ACE2
|
|||
|
Orf3a |
Increased formation of DMVs |
Electron Microscopy |
|
|
|
Electron Microscopy |
Apolipoprotein A-I |
|||
|
Orf3b |
Downregulates IFN-I ( Human type I interferons) signaling |
- |
|
|
|
Orf6 |
Interferon antagonist |
- |
|
|
|
Orf7a |
Favors apoptotic activity of infected cells |
Diffraction |
|
|
|
Orf7b |
Favors apoptotic activity of infected cells |
- |
|
|
|
Orf8 |
Interferes with immune responses |
X-Ray Diffraction |
|
|
|
X-Ray Diffraction |
|
|||
|
Orf9b |
Interferon antagonist |
X-Ray Diffraction |
|
|
|
X-Ray Diffraction |
|
|||
|
Electron Microscopy |
Human mitochondrial receptor Tom70 |
|||
|
Orf9c |
|
- |
|
|
|
Orf10 |
|
- |
|
|
|
E |
Envelope (E) protein, involved in virus packaging |
Solid-State NMR |
|
|
|
M |
Membrane (M) protein, involved in virus packaging |
- |
|
|
|
N |
Nucleocapsid (N) protein, involved in RNA binding and
packaging |
X-Ray Diffraction |
|
|
|
X-Ray Diffraction |
|
|||
|
X-Ray Diffraction |
monoclinic crystal form |
|||
|
X-Ray Diffraction |
|
|||
|
X-Ray Diffraction |
|
|||
|
X-Ray Diffraction |
|
|||
|
NMR |
|
|||
|
X-Ray Diffraction |
|
|||
|
X-Ray Diffraction |
|
|||
|
NMR |
7mer
dsRNA
|
|||
|
NMR |
10mer
ssRNA
|
|||
|
X-Ray Diffraction |
|
|||
|
X-Ray Diffraction |
|
|||
|
X-Ray Diffraction |
|
|||
|
X-Ray Diffraction |
|
|||
|
|
||||
|
Results
of the project “Prediction of the structure of SARS-CoV-2 proteins with
artificial intelligence techniques”, funded by Axencia Galega de Innovación
(GAIN) and co-funded by FEDER, and supported by Consellería de Economía,
Emprego e Industria, reference: IN845D-02. Resultados do
proxecto “Predición da estrutura de proteínas de SARS-CoV-2 con técnicas de
intelixencia artificial”, subvencionado pola Axencia Galega de Innovación
(GAIN) e cofinanciado con cargo aos fondos FEDER, e apoiado pola Consellería
de Economía, Emprego e Industria, referencia: IN845D-02. |
||||